R is undeniably a must-use language. Especially for data visualization. But R can sometimes be a little bit slow when dealing with big datasets. If you don’t need to create awesome graphs or don’t have time to wait, there’s an alternative in Python that can be quite fast for data manipulation. The Python Data Analysis Library, pandas, provides an easy way to manipulate data in python.
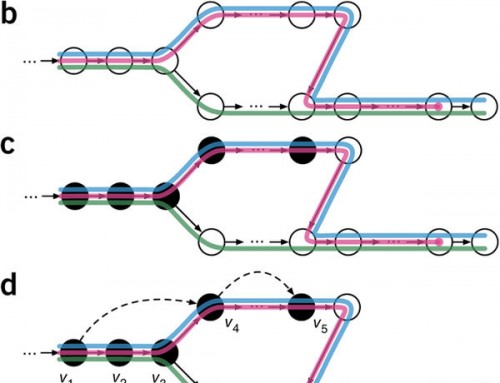
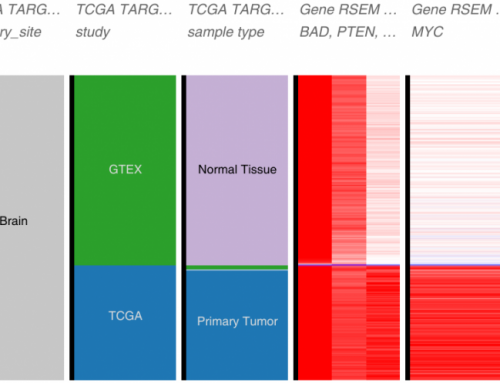
Recently, I had to deal with a big gene expression file (21024 genes x 3081 samples) derived from The Cancer Genome Atlas.
The RPKM were log-tranformed and I wanted to get back the untransformed (raw) data. The transformation applied to the data was : log10 (1000* RPKM +1). I was in a hurry so I decided to use pandas instead of R. :
Pandas offers two main data structures : Series and Dataframe. Series can be compared to R’s vectors and Dataframe can be compared to, well, R’s dataframe.
The package allows indexing and manipulation of the data and provides several functions whose purpose is to make our life easier. For example, dataframes can be created directly from dictionaries.
I’ve listed some basic functions implemented in pandas and their equivalent in R below.
| With pandas | With R | Description |
|---|---|---|
df.head() |
head(df) |
Display first lines |
df.tail() |
tail(df) |
Display last lines |
df.describe() |
summary(df) |
Get statistics |
df.shape() |
dim(df) |
Get dimensions |
df |
df[df$col=='valueForRow',] |
Retrieve a given row |
df.ix[0:2,1] |
df[0:2,1] |
Slicing |
df.index |
rownames(df) |
Get row names |
df.columns |
colnames(df) |
Get column names |
For my gene expression data situation, this is what I did:
import pandas as pd
df = pd.read_csv('TCGA.txt', sep='\t', dtype='object', header=0)
raw = (10**df[range(1,len(df.columns))].astype('float64')-1)/1000
raw_col = raw.columns
raw['Gene'] = df['Gene']
raw = raw[['Gene'] + list(raw_col)]
raw.to_csv('TCGA.raw.txt', sep='\t')
Loading the data on my computer took about 22.6 seconds, transforming it, 14.6 seconds and writing it back to a text file, 45.1 seconds.
You'll find more information in the pandas documentation . Have fun !



Pretty impressive speed!
Especially if you consider that the equivalent awk script runs in as much as 70 seconds:
awk ‘{for (i=1; i<=NF; i++) printf("%.2f\t", ((10**$i)-1)/1000); printf("\n")}'