The first thing I have learned in R is how to load a table. Usually, when you start your R journey, someone more knowledgeable will tell you how to do this very first action. It will typically be:
data<-read.table("~/SomeFolder/datafile.txt")
You probably will be adding various parameters into the brackets such as “row.names=0” or “header=TRUE” or, “sep="\t"“, to make sure you are reading your file correctly.
And this is perfectly fine, as a loading method of small datasets.
However, to maximize your time, let’s go over the common problems and, more importantly, their solutions new users can encounter when loading a dataset into R.
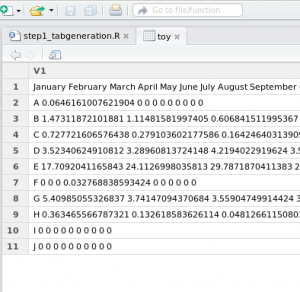
Problem #1: All the data is loaded into one column
Solution: This is fairly common. You need to specify the separator!
This can be achieved by adding sep=”XXX” into the read.table() function. In the place of XXX, you will put in your separator. Typically, it can be either the semi-column character (;) or a comma (,)
Problem #2: R didn’t understand I needed column names and rownames and put them into the data
Solution: This is rare. But you do need to specify if you table has a header and rownames!
This can be achieved by adding header=TRUE and rownames=0 into the read.table() function.
Problem #3: My table loaded fine but the data is set to factors. I don’t understand why!
> data<-read.table("data.txt")
> class(data[,1])
[1] "factor"
Solution: Tell R you don’t want factors! The as.is parameter can help you do that! Note about factors: Although sometimes very useful, this datatype can be very hard to deal with, especially if you want to do calculations on numeric values in your data frame. When using the read.table function, R’s default is set to convert all variables into factors.
> data<-read.table("data.txt",as.is = TRUE)
> class(data[,1])
[1] "character" Problem #4: My table takes a long time to load!
R is efficient at loading data only when it knows in advance what you are loading. As seen in problem #3, R will automatically convert all string data to factors. This means that the conversion is part of the loading step.
Solution: Exactly as seen in problem #3, tell R you don’t want factors!
Not convinced? Here is a performance review:
data.txt contains a RNASeq dataset: 21024 genes for 3080 patients.
> data<-read.table("data.txt",header=TRUE,sep="\t")
> elapsed
771.274 seconds
> data<-read.table("data.txt",header=TRUE,as.is=TRUE,sep="\t")
> elapsed
184.764 seconds
This is a 4 times faster load! You could cut your loading time even more, if you save your loaded table in a RData format (rda).
> load("data.rda")
> elapsed
35.915 seconds
But of course, loading a RData file implies loading it once into R…
I encourage you to explore other options the read.table() function offers, you might be surprised!
A side note about read.delim(): this function is a wrapper for read.table(), with only the most common features in it. Use at your own risk 🙂


I don’t understand your side note. What is the “danger” with read.delim? I always use it for my .txt files. Can you explain what you meant? Thanks!! 😀