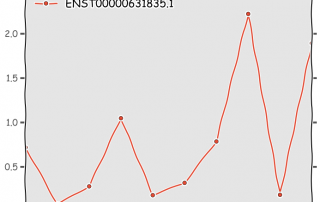
Understanding how kallisto works
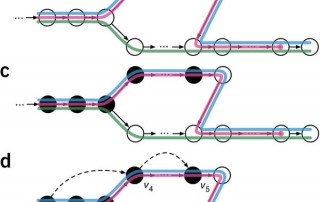
In 2016, Bray et al. introduced a new k-mer based method to estimate isoform abundance from RNA-Seq data. Their method, called kallisto, provided a significant improvement in speed and memory usage compared to the previously used methods while yielding similar accuracy. In fact, kallisto is able to quantify expression in a matter of minutes instead of hours. Since it is so light and convenient, kallisto is now often used to quantify expression in the form of TPM. But how does [...]