Overfitting and Regularization
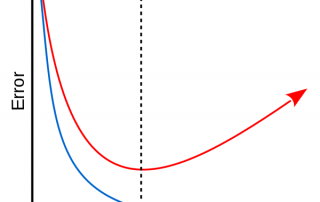
This series of articles on machine learning wouldn't be complete without dipping our toes in overfitting and regularization. Overfitting The Achille's heel of machine learning is overfitting. As machine learning techniques get more and more powerful (large number of parameters), exposure to overfitting increases. In the context of an overfit, the model violates Occam's razor's principle by generating a model so complex that it begins to memorise small, unimportant details (with no true link to our target) of the training set. [...]